 Qiime2 classification validator
Qiime2 classification validator

The recipe generates simulated sequencing reads from NCBI accession numbers. It then performs classification with a machine-learning based tool available in Qiime2.
In the third step evaluates and reports the accuracy (the percent of reads were classified correctly) of the classification.
The input to the recipe is a list of NCBI accession numbers:
AP012081
AP011279
DQ288268
AP014537
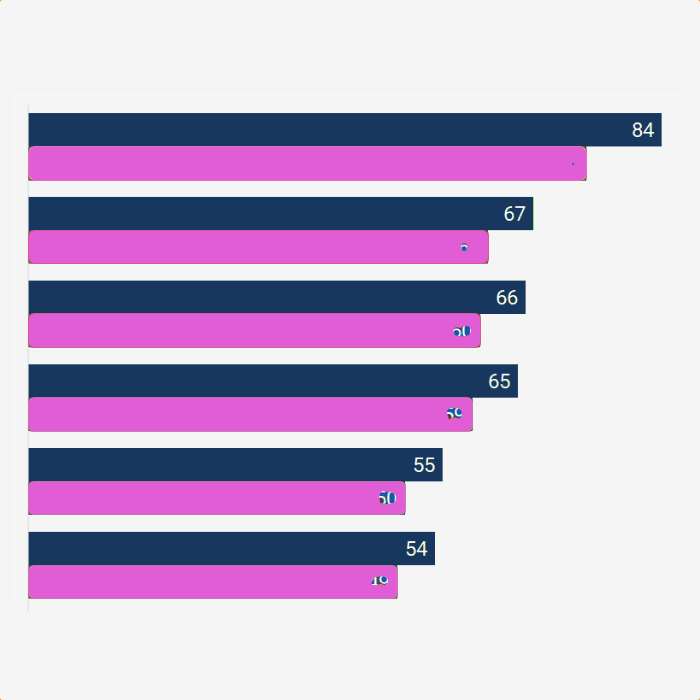
The output is a table with a taxonomic assignment of each read.
accession expect actual percent taxid title
MG570454 1000 25 2.5 90988 Pimephales promelas isolate NEFC F17-076 mitochondrion, complete genome
MG570425 1000 40 4.0 28800 Notemigonus crysoleucas isolate NEFC F16-041 mitochondrion, complete genome
MG570424 1000 48 4.8 7971 Catostomus commersonii isolate NEFC F16-032 mitochondrion, complete genome
MG570419 1000 48 4.8 67558 Semotilus atromaculatus voucher NEFC F16-365 mitochondrion, complete genome
MG570417 1000 29 2.9 407093 Rhinichthys obtusus voucher NEFC F16-301 mitochondrion, complete genome
Copy recipe
You need write access to the
original recipe
to edit.
Recipe Interface Builder
Click the buttons on the right to create new fields.
