 Bioinformatics Recipe Cookbook
Bioinformatics Recipe Cookbook
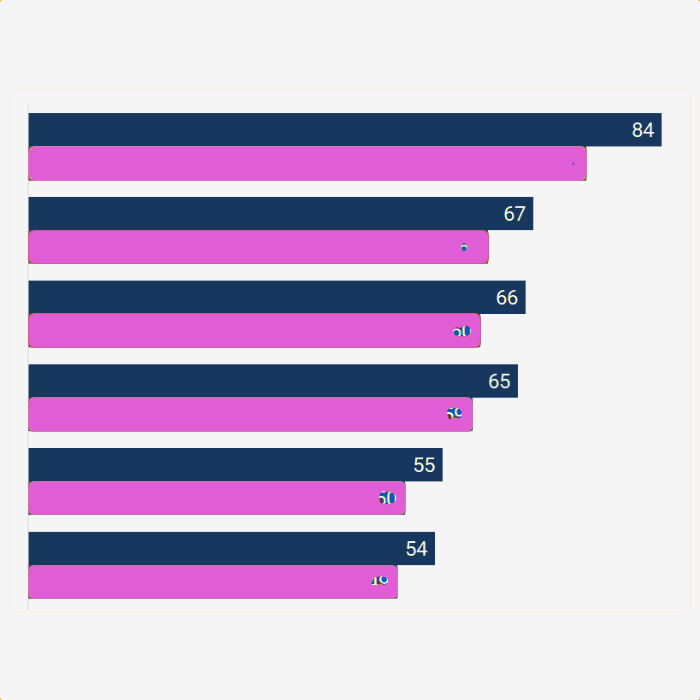
Results for Qiime2 classification validator
Completed
•
Runtime 1 minutes
•
updated 3.9 years ago
by
Istvan Albert
Results generated by running the recipe.
Parameters used during the run:
- Accession numbers:MISSING
File List
Files created by the recipe run:
classification_qiime2.txt
4.3 KB
code/simulate.py
2.3 KB
code/validate.py
3.9 KB
input/accessions.txt
638 bytes
input/reads.fq
13.4 MB
input/reads_manifest.txt
106 bytes
input/reference.fa
807.5 KB
input/taxa_lineage.tsv
5.7 KB
input/taxonomy.txt
5.2 KB
output/classified.qza
1.8 MB
output/classified.tsv
5.1 MB
output/feature-table.biom
3.1 MB
output/feature_table.qza
1.4 MB
output/feature_table.tsv
2.2 MB
output/reads.qza
1.5 MB
output/reference.qza
251.0 KB
output/species_table.qza
42.3 KB
output/taxa_lineage.qza
6.1 KB
output/trained_reference.qza
506.2 KB
recipe.sh
6.0 KB
runlog/input.json
1.3 KB
runlog/stderr.txt
997 bytes
runlog/stdout.txt
974 bytes
Output Messages
Messages printed to the standard output stream:
Imported input/reference.fa as DNASequencesDirectoryFormat to output/reference.qza Imported input/taxa_lineage.tsv as TSVTaxonomyDirectoryFormat to output/taxa_lineage.qza Saved TaxonomicClassifier to: output/trained_reference.qza Imported input/reads_manifest.txt as SingleEndFastqManifestPhred33V2 to output/reads.qza Saved FeatureTable[Frequency] to: output/feature_table.qza Saved FeatureData[Sequence] to: output/represetative_seqs.qza Exported output/represetative_seqs.qza as DNASequencesDirectoryFormat to directory output Saved FeatureData[Taxonomy] to: output/classified.qza Exported output/feature_table.qza as BIOMV210DirFmt to directory output Exported output/classified.qza as TSVTaxonomyDirectoryFormat to directory output Saved FeatureTable[Frequency] to: output/species_table.qza Exported output/species_table.qza as BIOMV210DirFmt to directory ./ Requirement already up-to-date: plac in /home/www/miniconda3/envs/qiime2/lib/python3.6/site-packages (1.2.0)
Other Messages
Messages printed to the standard error stream:
WARNING: pip is being invoked by an old script wrapper. This will fail in a future version of pip.
Please see https://github.com/pypa/pip/issues/5599 for advice on fixing the underlying issue.
To avoid this problem you can invoke Python with '-m pip' instead of running pip directly.
% Total % Received % Xferd Average Speed Time Time Time Current
Dload Upload Total Spent Left Speed
0 0 0 0 0 0 0 0 --:--:-- --:--:-- --:--:-- 0
0 0 0 0 0 0 0 0 --:--:-- --:--:-- --:--:-- 0
% Total % Received % Xferd Average Speed Time Time Time Current % Total % Received % Xferd Average Speed Time Time Time Current
Dload Upload Total Spent Left Speed
0 0 0 0 0 0 0 0 --:--:-- --:--:-- --:--:-- 0
100 3992 100 3992 0 0 125k 0 --:--:-- --:--:-- --:--:-- 125k
